Numerous studies have linked suppression of specific regulatory and signaling proteins to cancer phenotypes and their associated gene expression changes. Identifying new causal links between mutations and disruptions of these proteins to cancer states, however, requires the integration of data at both the expression and signaling/regulatory protein level. We have implemented a software system to discover new causal subnetworks from mixed phospho-proteomic, mutation and expression data. By presenting the subnetwork discovery requirement as both a constrained optimization problem and a task for Bayesian inference, we are able to apply several algorithms and combine their results in a meaningful way.
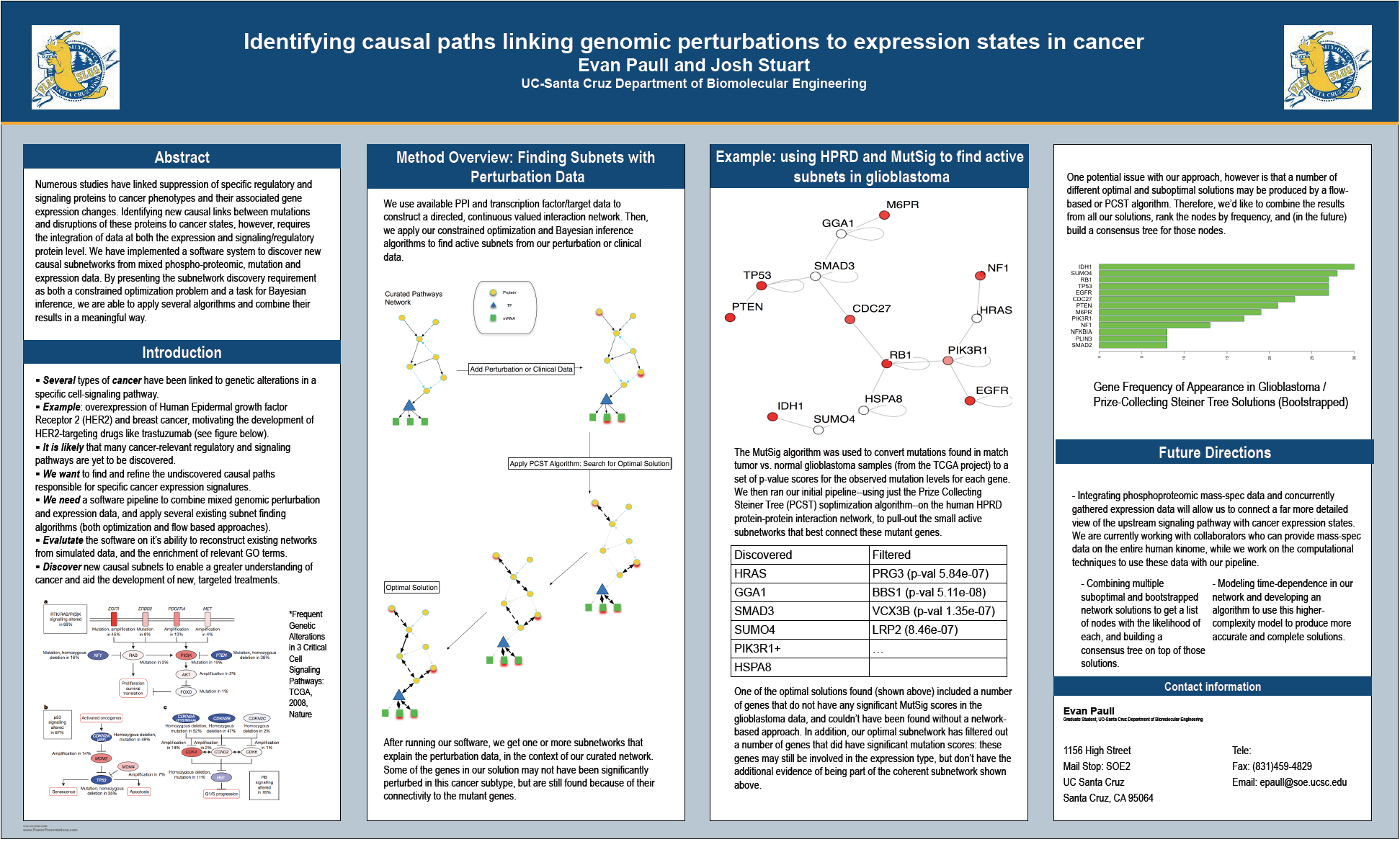
The MutSig algorithm was used to convert mutations found in match tumor vs. normal glioblastoma samples (from the TCGA project) to a set of p-value scores for the observed mutation levels for each gene. We then ran our initial pipeline--using just the Prize Collecting Steiner Tree optimization algorithm--on the human HPRD protein-protein interaction network, to pull-out the small active subnetworks that best connect these mutant genes. We show how our approach may be used on mutation data to discover new genes that are involved in cancer phenotypes, even when those genes don't have a significant mutation rate, and discuss the future applications to cancer.
Poster: